Cell-free systems are mainly used for in-vitro protein synthesis and can be considered as a powerful tool in genetic code reprogramming, involving the amber codon. In this use, the non-proteinogenic amino-acids are incorporated into proteins by charging them to suppressor-tRNAs molecules that reprogram the existing codons. However, cell-free systems are also used to engineer genetic circuits with applications for in-vitro biology or metabolic engineering.
Firstly, cell-free systems are being increasingly used as prototyping tools in synthetic biology since their use accelerates implementation into living cells (Chiao et al., 2016). Indeed, engineering cycles from design and construction to testing are faster in cell-free systems, taking less than 8 hours per cycle, compared to E. coli bacteria systems that take many days (Sun et al., 2014). In addition to speed, cell-free systems have the advantage to act as “biomolecular breadboard” since they preserve endogenous E. coli transcription-translation mechanisms (Sun et al., 2013).
Finally, after testing in cell-free systems, the engineered genetic circuits are implemented in-vivo. For example, constitutive DNA regulatory elements, such as sigma70 promoters and RBS, were shown to have the same relative effects on the strength of transcription and translation in E. coli and cell-free systems (Chappel et al., 2013). Cell-free systems were also used for rapid prototyping of 1, 4-butanediol synthesis (Wu et al., 2017).
Cell-free systems can be an interesting and efficient solution for in vitro biology or metabolic engineering.
In addition, cell-free systems can be used to synthesize fine-chemicals (Dudley et al., 2015). In one example, cell extract from a tpi knockout E. coli strain was used to convert cheap glucose (GLC) to dihydroxyacetone phosphate (DHAP). One equivalent of DHAP was produced per equivalent of GLC. The absence of triose-phosphate isomerase (tpi) enabled accumulation of DHAP and the absence of AMPnucleosidase (amn) prevented degradation of the catalytic ADP and ATP. Finally, DHAP was converted into an unnatural monosaccharide 5, 6, 7-trideoxy-D-threoheptulose-1-phosphate (TDHP) using E. coli endogenous aldolase FbaA (Bujara et al., 2010).
Finally, cell-free systems have been used to synthesize pharmaceutical natural products, such as polyketides (PKs). Nonribosomal peptides (NRPs) have been synthesized using cell-free systems as an alternative to extraction or chemical synthesis. For example, erythromycin (PK), doxycycline (PK) and daptomycin (NRP) are important antibiotics. To illustrate the efficiency of cell-free systems, 12 mg/L of D-Phe-L-Pro diketopiperazine were produced, using a cell-free system encoding for GrsA and GrsB1 enzymes that are necessary to the formation of DKP (Li et al., 2018).
In conclusion, the flexibility offered by the open nature of cell-free systems makes it a powerful platform for the production of in-vitro proteins, but also for fine chemicals, pharmaceutical natural products or for prototyping tools in synthetic biology.
Examples of cell-free system applications
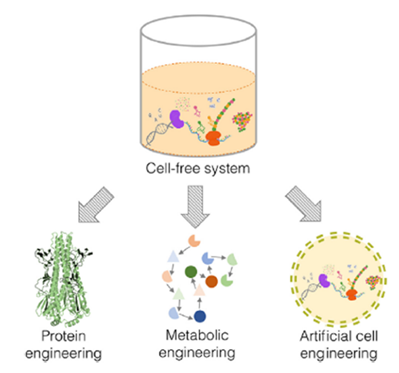
Source: Yuan Lu: Cell-free synthetic biology: Engineering in an open world, Kei Ai, Synthetic and Systems Biotechnology 2 (2017) 23-27.
Authors & Sources:
Bujara M., Schumperli M., Billerbeck S., Heinemann M., Panke S. 2010. Exploiting cell-free systems: implementation and debugging of a system of biotransformations. Biotechnology and Bioengineering 106:376–389.
Chappell J., Jensen K., Freemont P.S. 2013. Validation of an entirely in vitro approach for rapid prototyping of DNA regulatory elements for synthetic biology. Nucleic Acids Research 41(5): 3471–3481.
Chiao A.C, Murray R.M., Sun Z.Z. Development of prokaryotic cell-free systems for synthetic biology. 2016. BioRxiv. 10.1101/048710.
Dudley Q.M., Karim A.S., Jewett M.C. 2015. Cell-Free Metabolic Engineering: Biomanufacturing beyond the cell. Biotechnol J 10(1): 69–82.
Li J., Zhang L., Liu W. 2018. Cell-free synthetic biology for in vitro biosynthesis of pharmaceutical natural products. Synthetic and Systems Biotechnology 3:83-89.
Sun Z.Z., Hayes C.A., Shin J., Caschera F., Murray R.M., Noireaux V. 2013. Protocols for Implementing an Escherichia Coli Based TX-TL Cell-Free Expression System for Synthetic Biology. Journal of Visualized Experiments e50762.
Sun Z.Z., Yeung E., Hayes C.A., Noireaux V., Murray R.M. 2014. Linear DNA for rapid prototyping of synthetic biological circuits in an Escherichia coli based TX-TL cell-free system. ACS Synthetic Biology 3(6):387-397.
Wu Y.Y., Sato H., Huang H., Culler S.J., Khandurina J., Nagarajan H., Yang T.H., Van Dien S., Murray R.M. 2017. System-level studies of a cell-free transcription-translation platform for metabolic engineering. BioRxiv 10.1101/017814.



